Overview
We are living through the genomic revolution—and have only just crossed the starting line. Genetic diagnoses are being made in hours instead of weeks, new companies are engineering organisms for manufacturing purposes, and scientists are designing gene therapy delivery vectors using artificial intelligence.
What’s different? How are these types of technology now possible?
Over the course of a new series of posts, I’m going to cover four of the most essential components of this revolution: Sequencing, Synthesis, Scale, and Software. In the first post, I provided an overview of the exponential cost decline in DNA sequencing. The Human Genome Project took over a decade to complete, and required nearly $3 billion dollars. In the two decades since, we have scaled sequencing technology to the point that we are sequencing millions of people—an even faster rate of progress than the digital revolution. This has opened up a wide range of scientific and commercial opportunities.
This post will focus on DNA Synthesis, which is the workhorse of modern biotechnology.
Synthesis
While it hasn’t declined in cost at the rate of DNA sequencing, DNA synthesis technology is crucially important. An enormous number of molecular technologies use DNA as a substrate. As a result of COVID, many people are now intimately familiar with Polymerase Chain Reaction (PCR) and its use as a diagnostic tool. This DNA copying reaction is also an essential component of most modern molecular biology protocols. PCR requires oligonucleotides—normally abbreviated to oligos—which are short strands of DNA that prime the reaction.
Beyond PCR, all of the next-generation sequencing technologies (NGS) described in the first post heavily rely on oligos. For example, the Illumina flow cells that enabled massively parallel sequencing are covered in oligos.
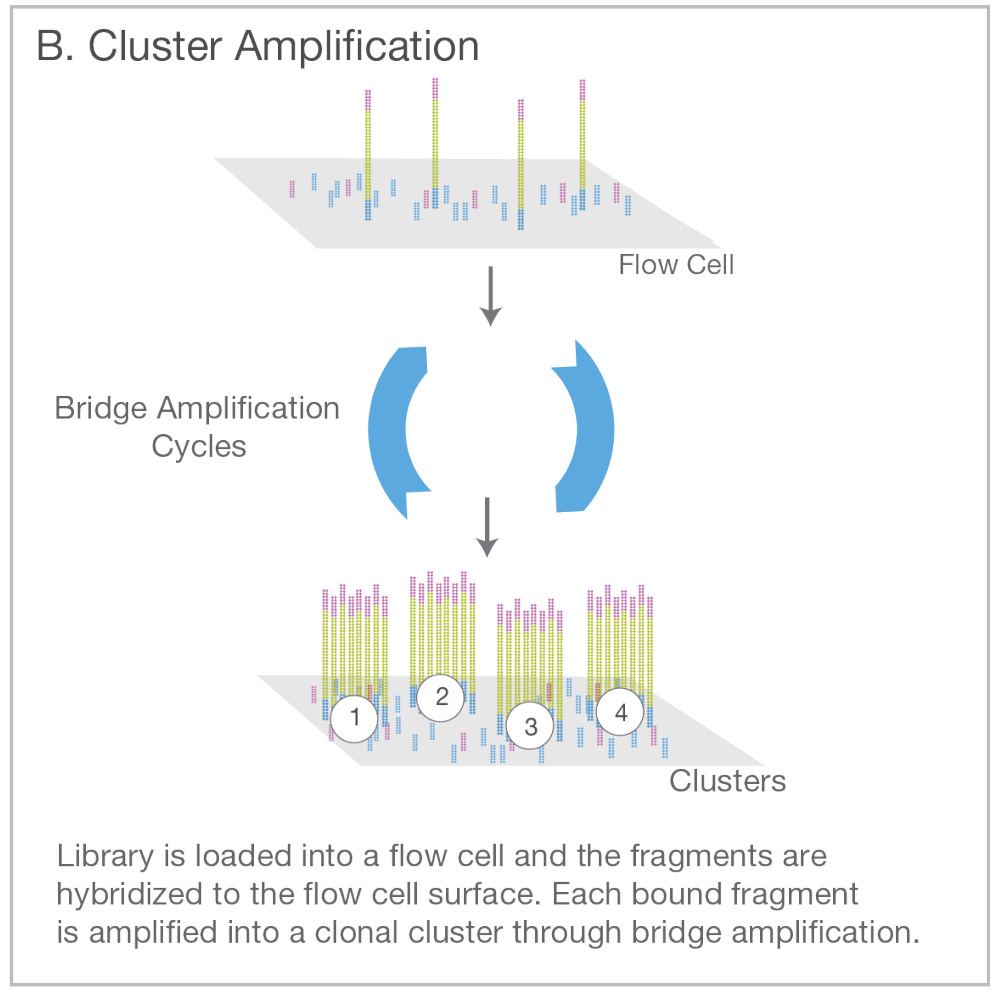
What about the breakthrough of CRISPR gene editing? Yep, also heavily reliant on oligos. The most revolutionary part about the CRISPR editing complex is that it is programmable. Short sequences of RNA called single guide RNAs (sgRNAs) determine the sequence that will be edited. sgRNAs are encoded in oligos.1
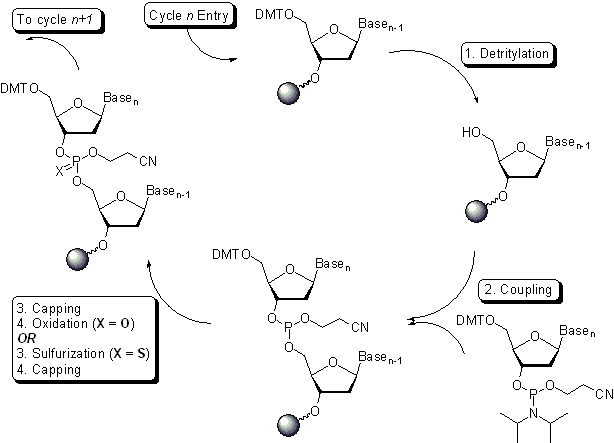
The major technical innovation has been the commoditization of oligo synthesis. These short sequences of DNA can be designed and ordered, with some providers offering turnaround times of a single day. Nearly all oligos are synthesized using phosphoramidite chemistry, which is a four-step cyclical reaction that was first developed in the 1980s.2
In the 1990s, a company called Affymetrix developed an array-based approach to oligo synthesis—with the goal of creating a measurement technology called DNA microarays. Here, the story mirrors that of next-generation sequencing—where extensive process engineering was applied to massively parallelizing one fundamental reaction.3
Since then, many different engineering approaches have been taken to synthesize large numbers of oligos simultaneously. Companies such as Integrated DNA Technologies (IDT) and Agilent provide this type of service, and Twist Bioscience has become one of the most popular providers of complex pools of oligos.
Beyond applications like PCR, NGS, and CRISPR, large-scale oligo synthesis has enabled totally new types of technologies to be developed. It is now possible to use NGS technologies to measure properties of individual cells. This is a crucial new capability. It has opened the door for projects like the Human Cell Atlas which aims “to create comprehensive reference maps of all human cells—the fundamental units of life—as a basis for both understanding human health and diagnosing, monitoring, and treating disease.”
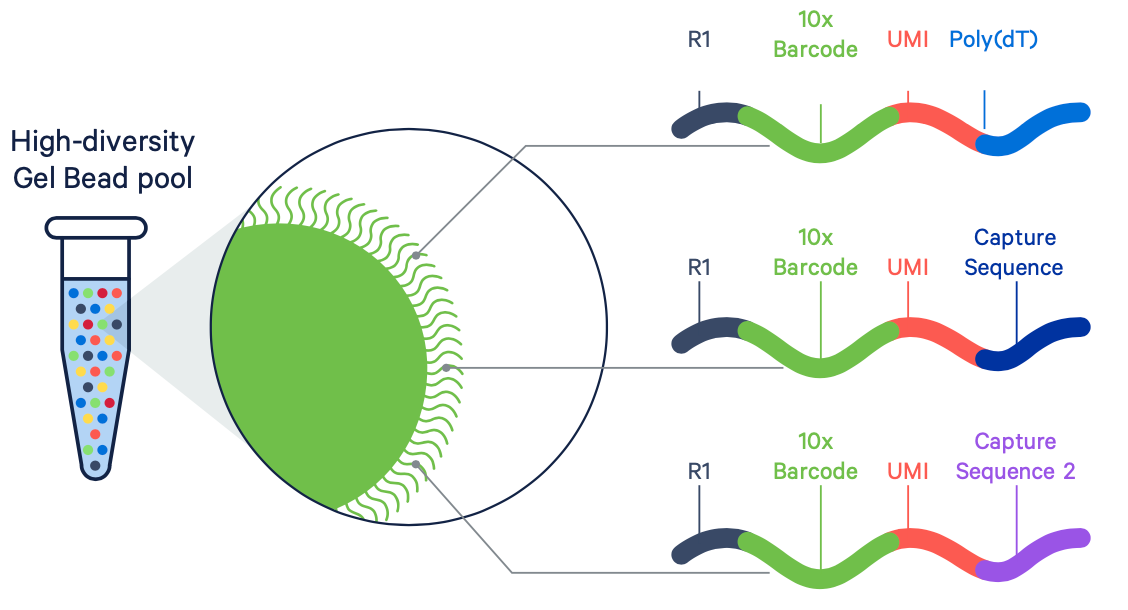
A company called 10X Genomics is the major provider of single-cell technologies. Their approach uses gel beads coated with oligos containing barcode sequences that can be used to capture and identify individual cells for measurement. This an example of a breakthrough technology built on top of both Sequencing and Synthesis.
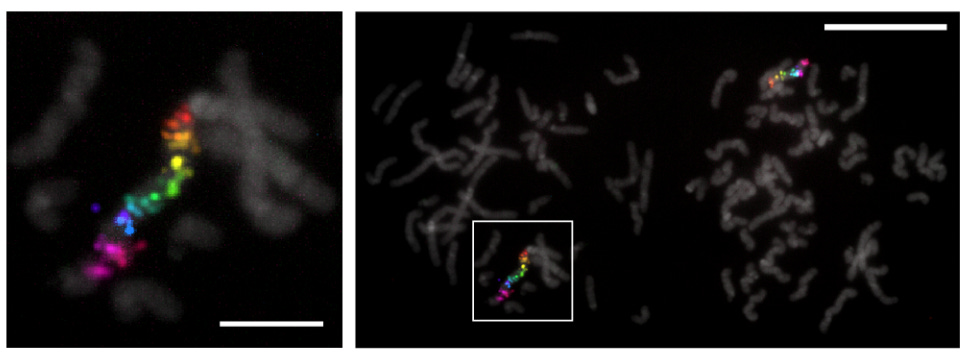
Another of my favorite examples is that we can now paint with synthetic DNA. I worked for two years in a lab developing this type of technology. The core idea is that it is possible to program oligos to target and visualize specific nucleic acid sequences in space. This makes it possible to resolve the 3D organization of the genome, and to image thousands of gene transcripts simultaneously.
The field of synthetic biology aims to expand our ability to use life as an engineering substrate. Synthetic DNA is essential for this pursuit. This explains why Gingko Bioworks—one of the largest synthetic biology companies—is also the largest consumer of DNA from Twist Bioscience. The company estimated that they ordered around one billion bases of DNA over the last four years.
Sequencing and Synthesis may also solve our looming digital storage bottleneck. As a species, we are generating and collecting data at a rate that will soon make efficient physical data storage a hard problem. DNA offers an enticing solution, seeing as it is the only highly stable nanoscale information storage technology that we know of. Conceptually, an exabyte of data could fit in the palm of your hand. Many labs and companies are working on combing large-scale synthesis and sequencing to reliably store information in DNA and retrieve it.
So what else might come next?
There is still room for an enormous amount of progress in DNA synthesis. The key axes for improvement are increasing sequence length and decreasing cost. Innovation in this area could lead to the holy grail of synthetic biology and genome engineering: the ability to routinely synthesize entire genes, chromosomes, and even genomes.
One of the major efforts to realize this dream is the GP-write project, which aims to “oversee a reduction in the costs of engineering and testing large genomes in cell lines more than 1,000-fold within ten years.” This is an incredibly ambitious collaboration between academic labs and several companies. Twist Bioscience is a member of this project, along with Ansa Biotechnologies4 and DNA Script, which are both startups working on next-generation enzymatic DNA synthesis technologies.

Twist Bioscience is also pursuing this direction, creating powerful market dynamics. Cost reduction for synthesis would have an enormous impact. Commoditized large-scale oligo synthesis is essential for foundational technologies like PCR, and has made it possible to develop new approaches for single-cell sequencing, complex spatial imaging, and even digital data storage. This set of examples represents a sparse sampling of a rich space of other applications—such as protein and metabolic engineering and DNA encoded libraries for drug discovery. What will become possible if we can routinely synthesize entire genomes at a reasonable cost?
At a first approximation, this could accelerate and refine the engineering process for several therapeutic modalities built on top of DNA—such as gene and cell-based therapies. But ultimately, therapeutics is a subset of biotechnology. This sentiment was expressed by many of the attendees of last week’s Built with Biology conference in Oakland.


In his keynote, Eric Schmidt said that he felt “your industry is at the same stage mine was in 40 years ago when I started my career in tech.” Just as it was with the Internet revolution, it’s difficult to speculate about what form this will take in practice—but my conviction is that we have barely started our exploration of where this will go.
Next time: Scale
The first post in this series provided an overview of the progress that has been made in Sequencing. My aim in this chapter has been to focus on advances in Synthesis, what it has enabled, and where the frontier is for this technology. Next time, the story will expand to describe the Scale of biological experimentation and engineering that is now possible.
There will be more in-depth coverage on the importance of this advance as this series of posts progresses.
If you want to know more, a really accessible blog post describing phosphoramidite chemistry can be found here.


