It's all too easy to dismiss the future. People confuse what's impossible today with what's impossible tomorrow.
- George Church
Overview
Exciting work often happens at intersections. Many of the pioneers of molecular biology were originally physicists. Bioinformatics requires interdisciplinary work combining computer science and biology. The modern fields of genomics and synthetic biology are melting pots of disciplines, combining the toolkits of molecular biology, bioinformatics, and engineering. One of the leading groups of this new antedisciplinary science is the Church Lab at Harvard1.
The core emphasis of the Church Lab over the years has been to attract talented scientists and engineers from a variety of disciplines, and to give them the time and space to collaborate on riskier projects that may not be pursued elsewhere. A large number of companies have resulted from the projects that succeeded. Examples include eGenesis, a company using CRISPR to engineer pig organs for safe human transplantation, and Kern Systems, a startup building systems to use synthetic DNA for highly efficient data storage.
In 2019, the Church Lab published a landmark study in Science describing a promising new strategy for designing viral vectors for gene therapy. The paper was the result of a four year effort to combine three exponentially improving technologies: 1) massively parallel DNA synthesis, 2) next-generation DNA sequencing, and 3) machine learning. Essentially: the ability to inexpensively “write” and “read” DNA molecules, and the computational tools to learn complex patterns in new data.
The team used these technologies to build a platform for comprehensively evaluating how changes in protein sequence impact the formation of Adeno-associated virus (AAV) capsids, one of the core vehicles for delivering gene therapies to cells. Several of the authors went on to found Dyno Therapeutics based on this work.
In an exciting 2021 study, members of Dyno in a collaboration with Google leveraged supervised machine learning2 to design new capsids, but it turns out that the researchers have also explored an unsupervised learning3 strategy for this problem. Recently Sam Sinai, Nina Jain, George Church, and Eric Kelsic posted a preprint entitled “Generative AAV capsid diversification by latent interpolation” describing this work.
Sam Sinai described the effort of putting this paper together in the following way:

I, for one, am certainly glad that the group shared this interesting work.
Key Advances
In order to understand the results from this study, it is important to think about the general flow of delivering a gene therapy using AAV as a vector:
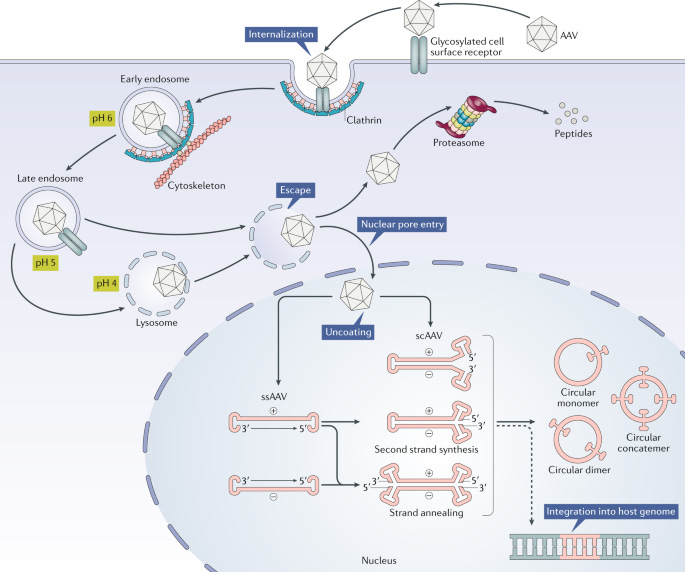
Gene therapies can be bundled into the icosahedral AAV capsid containers which can deliver the payload to the cell where it is ultimately integrated into the host genome. While this is a compelling delivery strategy, there are certain aspects of naturally evolved capsids that make them suboptimal vehicles. One of the crucial problems is that the human immune system is trained throughout life to detect and prevent viral infection. In order to improve delivery efficiency, it would be ideal to be able to engineer capsids that could better evade the immune system.
One of the challenges in capsid engineering is that due to the complexity of the protein structure, “introducing multiple mutations often breaks capsid assembly.” To circumvent this, the 2021 study combined DNA synthesis and sequencing to enable massively multiplexed measurement of the viability of sequence changes to AAV capsid production. The experimentation was guided by an ML strategy called active learning to iteratively generate new sequences. This powerful strategy enabled the team to discover a diverse array of new capsid sequences with novel properties.
While this strategy is powerful and producing promising results, it doesn’t incorporate evolutionary information into the model. Beginning as early as 2017, Sinai has been exploring the use of a Variational Auto-Encoder (VAE) to model the latent sequence space of AAV capsids to incorporate evolutionary and experimental information. What does this mean?
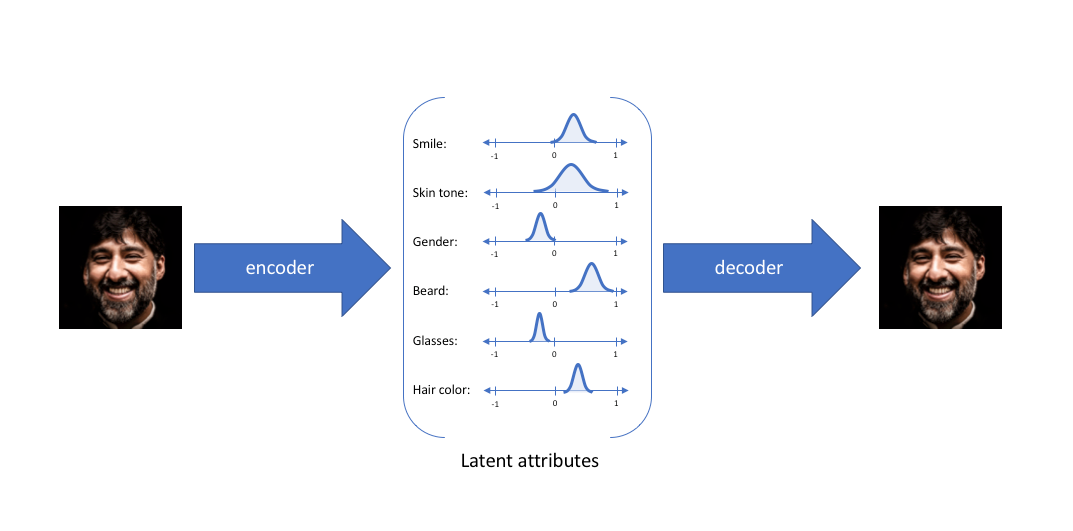
To build intuition for VAEs, let’s think about faces4. In a given set of faces, there are certain characteristics of each face that distinguish it from the others. At a conceptual level, these are the latent attributes of that face:
In this depiction, we see that in a VAE, latent attributes are represented as probability distributions. The encoder and decoder are both neural network models that are trained to optimize the accuracy of the data constructed from the latent attributes. By using probability distributions to represent latent attributes, VAEs must learn a smooth representation of the latent space of the data, in order to be able to accurately reconstruct the output for any given input.
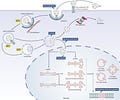
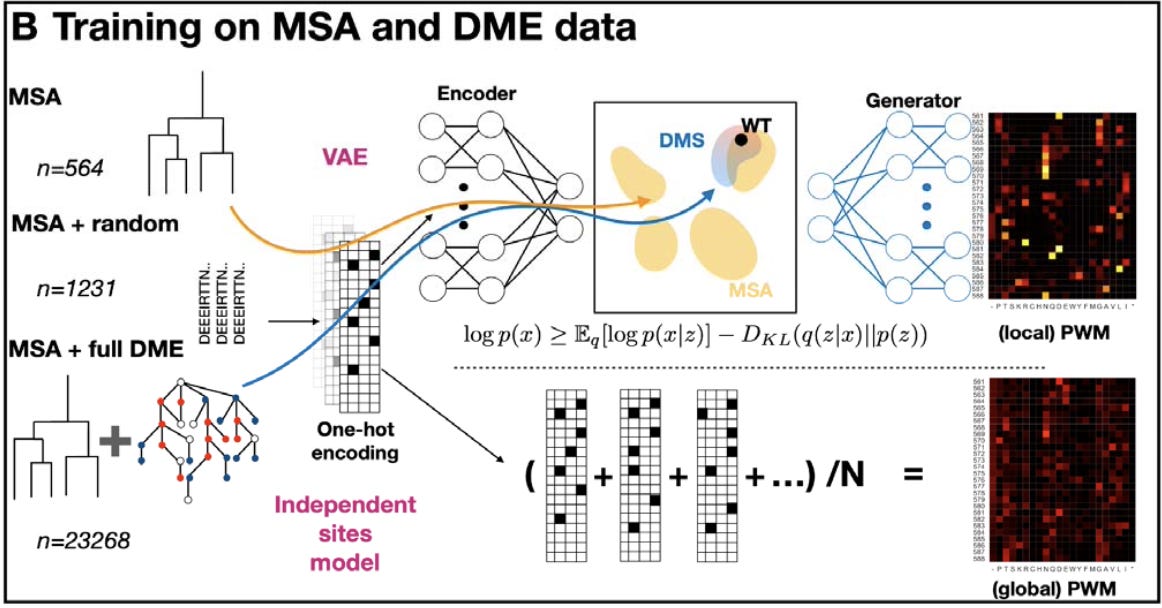
In this work, we are thinking about AAV capsids instead of faces. The models were learning latent representations of sequence space based on evolutionary information (multiple sequence alignment data of protein sequences) and empirical information (the training data from the study described earlier) describing AAV capsids:
With a learned representation of the latent space, they explored it in the search for new and improved capsids:
The latent space provides a way to efficiently search through AAV sequence space for variants with diverse characteristics.
Results
The authors established three bins for their generated results:
Viral variants likely arising from dysfunctional genes (observed in production due to low frequency cross-packaging into viable capsids) termed non-viable, viral variants likely capable of generating functional assemblies termed viable, and viral variants that are present at very high counts, termed viable+
This study evaluated a baseline model (an independent-sites protein model) and a simple VAE architecture on three datasets: 1) MSA alone (MSA), 2) MSA supplemented with a randomly selected subset of 667 viable variants (MSAr), and 3) MSA supplemented with the 22,704 viable variants 1-21 mutations away from wildtype (MSA+)5.

The results were quite promising:
Encouragingly, we found more than 100 viable variants with 25 or more substitutions in the 28- aa region, a majority of which came from the VAE designs. Of the top 10 variants produced, 9 were proposed by VAEs, and 1 by IS-MSA+. Of the top 100 scoring variants produced, 70 were proposed by the three VAE models, and the rest were proposed by IS-MSA+
This is really noteworthy, because “most random changes in the amino-acid space within this region result in a non-functional capsid.” The complexity of the problem is what makes the results from this fairly simple unsupervised model so exciting.
There is certainly a lot more to explore in this paper, including some of the interesting differences in behavior between the baseline model and the VAE, but I will end my brief highlight of the results here.
Final Thoughts
The research and engineering coming out of Dyno Therapeutics is an example of the promise of combining rapidly improving molecular technologies with new ideas and tools from computing and machine learning. Because each of the core technologies in their platform (DNA synthesis, DNA sequencing, ML) is improving at lightning speed, it seems like each new publication demonstrates a considerable leap forward for AAV engineering.
I will note that if reading about advanced viral engineering during a global pandemic makes you feel a bit uneasy, you aren’t alone. The author and entrepreneur Rob Reid has been warning about the potentially civilization ending ramifications of engineered pandemics well before the entire world was grappling with COVID-196. As genomics and synthetic biology accelerate, it becomes increasingly possible for bio-terrorists to inflict damage at a scale that is an existential risk to society.
In other words, we are truly living through The Century of Biology. The same new knowledge and tools that offer practically limitless possibilities for our ability to live healthy and meaningful lives in partnership with Nature also introduce some of the most terrifying threats to our continued existence.
As we have collectively learned with COVID-19, our global genomic infrastructure is inadequate. However, we are only limited by our imagination and ability to collaborate effectively to enact change. Brilliant new strategies like detecting pathogens from wastewater and the breakthrough progress with mRNA vaccines show that these problems are not intractable. As genomics and synthetic biology continue to advance, establishing this infrastructure becomes both more achievable and more urgently necessary.
Thank you for reading this highlight of “Generative AAV capsid diversification by latent interpolation”.
If you’ve enjoyed reading this and would be interested in getting a highlight of a new open-access paper in your inbox each Sunday, you should consider subscribing:
That’s all for this week, have a great Sunday!
A lot has been said and written about George Church and his lab. This profile from 2016 provides a fun and concise overview, although there is certainly more that has happened since then.
Supervised ML learns the function that maps an input to an output based on examples of input-output relationships.
On the other hand, unsupervised learning aims to learn useful representations of unlabeled datasets.
The following choice of example and graphic come from this excellent post on variational auto-encoders. I would personally recommend subscribing to Jeremy Jordan’s posts if you are learning more about machine learning.
The empirical data here is a subset of the training data from the 2021 supervised learning study.
In this scenario, we literally wouldn’t be around to regretfully rewatch this talk in the way that many rewatched Bill Gates’s 2014 TED talk warning about our inadequate preparation for viral outbreaks as many did during COVID-19.