Designing regulatory DNA
Using generative ML to create DNA sequences that control gene expression
Engineers turn dreams into reality.
(from The Wind Rises, by Hayao Miyazaki)
Overview
It is increasingly hard to argue against the broad utility of machine learning for tackling challenging problems in the life sciences. This was demonstrated on July 15th, where the AlphaFold paper from DeepMind was published in Nature and the RoseTTAFold paper from the Baker Lab was published in Science on the same day.1
These papers both represent the enormous progress that powerful deep learning approaches have brought to the difficult problem of protein structure prediction. It’s hard to appropriately emphasize how exciting this is, but the pioneering Harvard computational biologist Debora Marks did a pretty good job:
As I’ve previously written about, deep learning is not only achieving state-of-the-art results in protein science, but is also leading to new breakthroughs in genomics. We are learning how to build increasingly powerful and flexible representations of the regulatory dynamics of genomes. Many scientists are also working on the important problem of building new tools to interpret the information that these models are capturing.
Understanding genome regulation is of fundamental scientific importance. Similarly, developing the ability to design new regulatory DNA is critical for the rapidly evolving engineering discipline of synthetic biology.
A new preprint from the Zelezniak Lab at Chalmers University of Technology in Sweden entitled “Supervised generative design of regulatory DNA for gene expression control” proposes a new method for this problem. The lead authors of this new study are Jan Zrimec and Xiaozhi Fu.
Key Advances
Genetics is an explanatory science. It is fundamentally concerned with understanding and explaining the relationship between genes and heredity. Born from the sequencing revolution, the discipline of genomics generalizes this to consider the entire relationship between genomes and biological variation.
Biotechnology, on the other hand, is not concerned with explanation. Biotechnology seeks to harness and repurpose the only programmable nanotechnology that we know of: living systems. One of the most successful branches of biotechnology has been genetic engineering. Genetic engineering was jumpstarted in 1973 with the development recombinant DNA technologies that spurred an enormous amount of research and commercial innovation, including the birth of Genentech.
While genetic engineering has been tremendously successful, the emerging discipline of synthetic biology seeks to leverage several powerful molecular technologies (recombinant DNA, DNA sequencing, DNA synthesis) and create new engineering abstractions to dramatically expand what we can build with biology. Drew Endy, one of the pioneers of this new discipline, puts it the following way:
Synthetic biology builds on genetic engineering by improving the workflow: DNA synthesis to decouple design and fab[rication], standards to enable coordination of labor, and abstraction to manage complexity.
Synthetic biology increasingly seeks to build entirely new genetic circuits to accomplish novel applications. One of the design bottlenecks is that it is challenging to effectively design the noncoding DNA that regulates the expression of the genes in the new sequences.
Here is how the authors of the paper describe this problem:
Multiple recent studies show that apart from tuning codon usage in gene coding regions, also the DNA sequence of non-coding regulatory regions must be fine-tuned in order to accurately control gene expression. Proper orchestration of gene expression depends on the interaction of regulatory patterns across the whole cis-regulatory structure around the gene, including promoters, terminators, coding and untranslated regions (UTRs).
The tools for engineering new regulatory DNA struggle to account for this complexity. Namely, approaches to the problem include randomly mutating the promoter sequence, or using more knowledge-guided approaches to combine multiple known sequence motifs. Machine learning has also been used to generatively design individual components, again focusing on promoter sequences. To move beyond this paradigm and improve the the ability to holistically design new regulatory DNA, the authors had an idea:
Based on recent achievements in modeling DNA and protein spaces, we hypothesize that state-of-the-art generative deep neural networks are capable of learning the entire DNA regulatory landscape directly from natural genomic sequences.
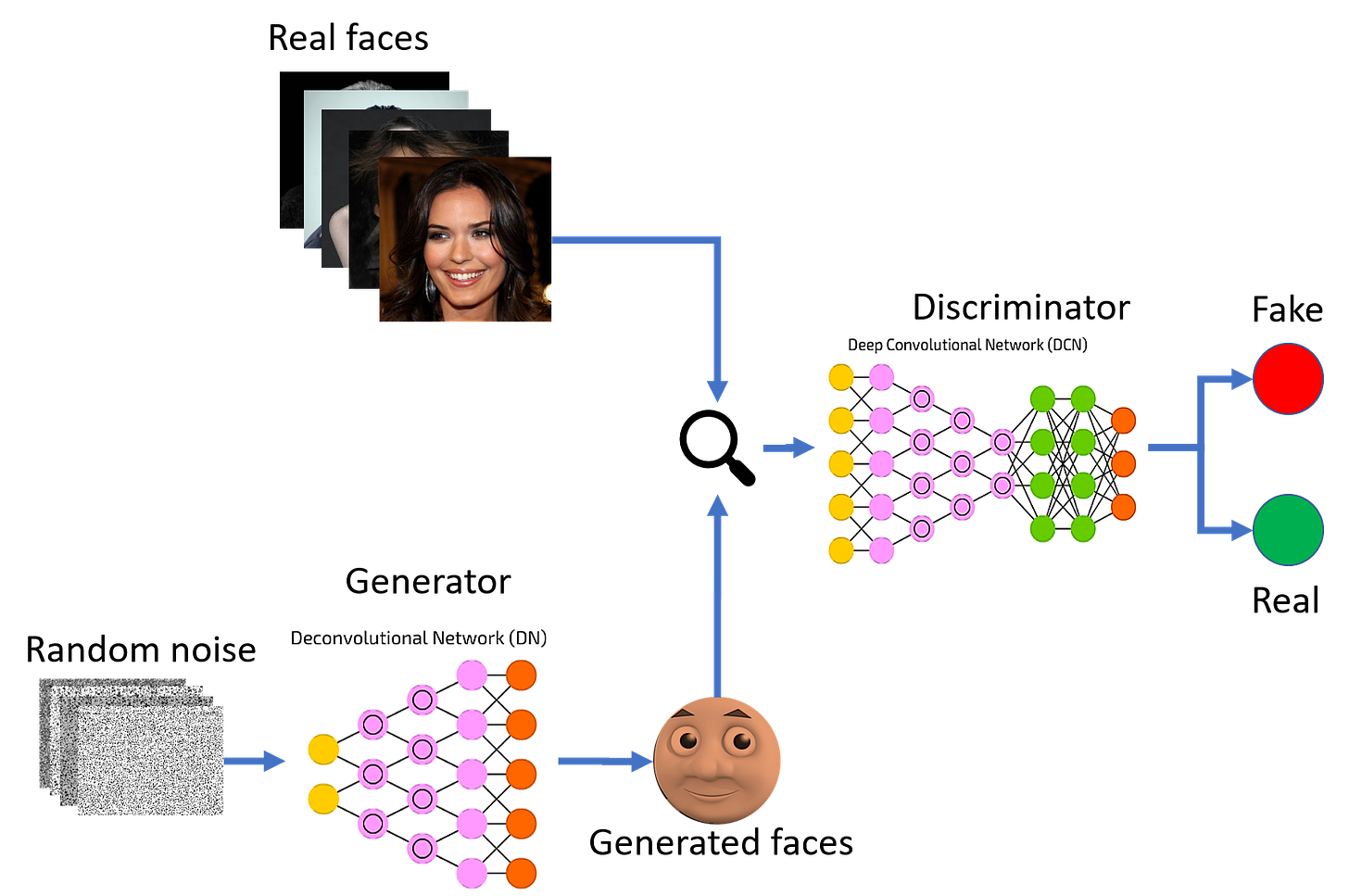
To do this, they built a new model called ExpressionGAN, which uses a type of machine learning model called a generative adversarial network (GAN). This type of model consists of two neural networks: the generator and the discriminator. The generator network is trained to generate sufficiently realistic data instances to make the discriminator network predict that it is real. The discriminator is trained to correctly classify between real and generated (fake) data. This schematic can be seen below for generating new images of faces:
GANs have generated absolutely incredible (bordering on scary) results across a wide variety of domains.2 For example, try playing the game Which Face is Real? which will very quickly drive home the message of how powerful these models are.
Here, instead of generating faces, the authors trained a GAN on natural genomic data, which “generates de novo functional regulatory DNA with prespecified expression levels.”
Results
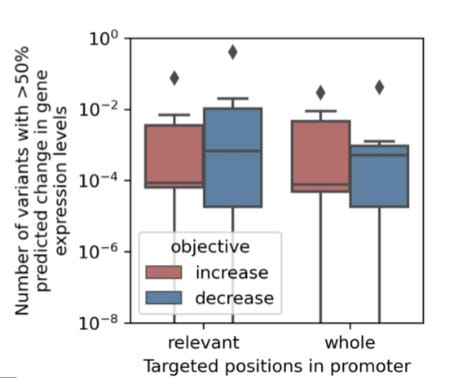
In order to evaluate their new model, the authors first carried out a random mutagenesis approach in yeast as a baseline. The mutagenesis strategy included an in silico screening step using a predictive model of gene expression. The results of this type of effort are modest:
When aiming to achieve an over 50% increase or decrease in mRNA expression levels, we found that on average, at most 0.3% of the sequence variants were predicted to achieve the desired effect when mutating 10% (40 bp) of whole promoter regions, which increased to 0.4% when mutating the most relevant promoter regions.
Next, they tested the predictions empirically, finding that “Of the tested variants, 40% corresponded with predictions, of which all were designed to decrease expression.”
This type of predictive performance means that designing regulatory DNA would require an iterative strategy, with multiple rounds of design and testing. With random mutagenesis coupled with in silico screening as a baseline, the authors moved to evaluate their GAN. One really interesting finding was that when they visualized the latent spaces of their model3, they found that it contained a distinct progression of ~6 orders of magnitude of gene expression:
But how well does this generative model perform?
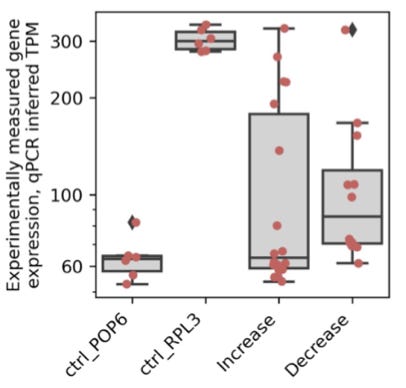
Overall, they “observed that experimental measurements of the mRNA levels produced by each construct achieved strong correlation with the predicted levels.” As can be seen from figure 2g, the model struggled a bit more with effectively generating sequences for the ~10 TPM4 range, but the model was overall quite effective, and had strong performance in the ~100 TPM range. Also, in terms of generating regulatory sequences for high gene expression, “4 out of 7 regulatory constructs (57%) displayed average expression levels that surpassed those of the natural RPL3 control by up to 2.7-fold.”
Overall, these empirical results and a wide variety of evaluation metrics used to characterize the generated sequences in the paper indicate that this is a promising research direction for the young engineering discipline of synthetic biology.
Final Thoughts
Synthetic biology is concerned with creating the tools and abstractions necessary to create increasing complex and powerful biological technology. Analogous to the process of connecting all of the computers on Earth to the Internet and the World Wide Web, it is nearly impossible to predict what this will lead to. One of the leading synthetic biology companies, Ginkgo Bioworks, has an entire creative and design department dedicated to exploring what the ramifications of synthetic biology might be. One of the members of this team named Eli Block shared a “mood board” of some of the designs they were using for inspiration:
The third photo in the second column is one of the most widely shared pieces of solarpunk art by an artist called Imperial Boy. To me, this represents the dream and promise of synthetic biology and biotechnology at large. Beautiful green cities, grown instead of built. Abundance, in alignment with our planet’s ecosystem.
In order to actualize this future, many engineers and scientists must continue to get up each morning and press forward on the frontier of engineering living systems. Each new insight and creation brings us a step closer. This preprint, which introduces a new method to harnesses the power of machine learning to generate regulatory DNA, represents an exciting step forward.
Thanks for reading this highlight of “Supervised generative design of regulatory DNA for gene expression control”. If you’ve enjoyed reading this and would be interested in getting a highlight of a new open-access paper in your inbox each Sunday, you should consider subscribing:
That’s all for this week, have a great Sunday! 🧬
Check out this incredible resource provided by DeepMind and the EBI: https://alphafold.ebi.ac.uk/
For a “deep” dive on Deepfakes, check out the new book “Deepfakes: The Coming Infocalypse” by Nina Schick.
I recently wrote a highlight of a visualization tool called VeloViz for single-cell genomics. Some of the background in that post could help with interpreting this figure.
TPM stands for Transcripts Per Kilobase Million.